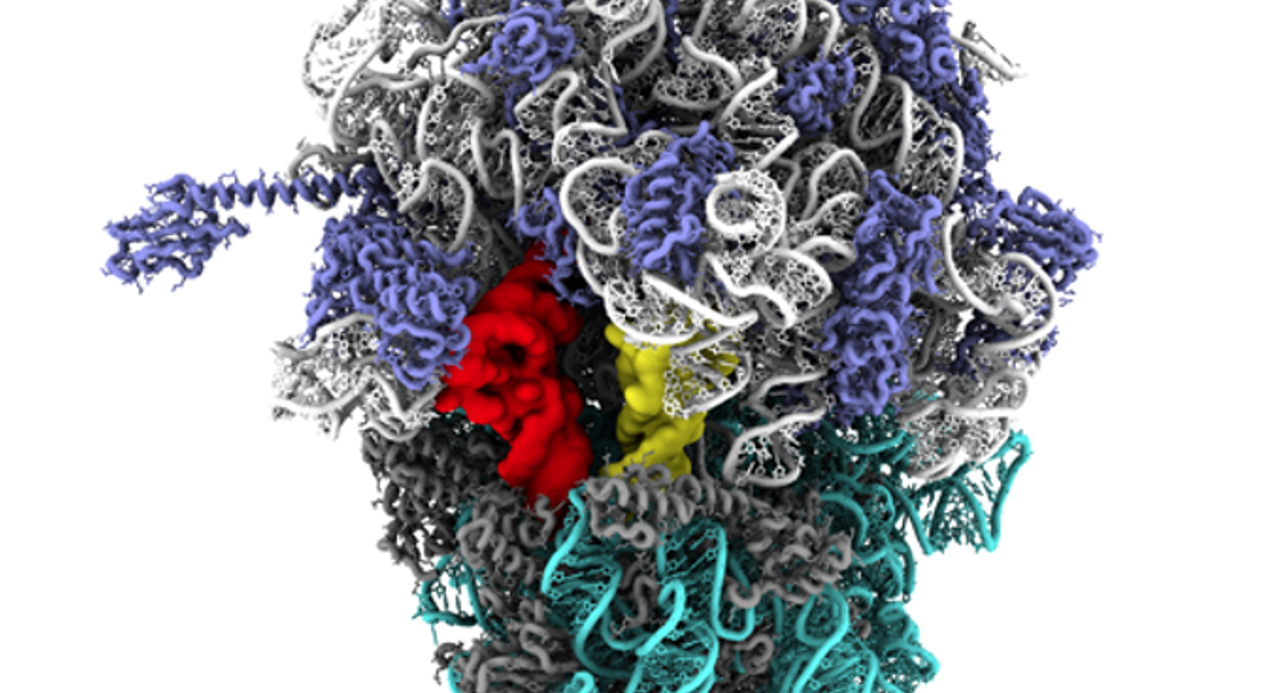
Northeastern's Prof. Paul Whitford uses the Discovery Cluster housed at MGHPCC to identify a novel motion within the ribosome.
Read this story in Northeastern's ITS blog
Earlier this year Nature Communications published its first research paper based on work performed exclusively on the Discovery Cluster at Northeastern University. Professor Paul Whitford from the Department of Physics at Northeastern, along with his graduate student Kien Nguyen, conducted research on the relationship between molecular structure and the dynamics in the ribosome, and authored Steric interactions lead to collective tilting motion in the ribosome during mRNA–tRNA translocation.
“Our study predicts how a precise movement in this nanoscale machine (the ribosome) directly controls biological function,” said Professor Whitford. “This opens up a wide range of medical applications that will aim to regulate the cell.”
According to Professor Whitford, since 2,000 researchers have been interested in how large-scale rotations in the ribosome enable biological function. But thanks to the high-end computing resources that Northeastern offers, his group’s work completes this picture by identifying a new rotation, as well as showing its regulatory role in the ribosome.
“Without Northeastern University’s research computing resources, this would not have been possible,” said Professor Whitford. “It is terrific how the Discovery Cluster allows us to explore and develop new ideas. This is exciting for me and my students.”
Discovery is the university’s shared high performance computing (HPC) cluster installed in the Massachusetts Green High Performance Computing Center (MGHPCC) facility and available for use to all Northeastern researchers and collaborators. A dedicated research computing team manages the cluster and provides a high level of support for Northeastern’s HPC researchers, including programming support, software installation and configuration, and algorithmic expertise in several research domains.
“The Research Computing team and I are proud to be able to support Paul and Kien in their research using Discovery that led to this achievement,” said Rajiv Shridhar, who directs Northeastern's Information and Technology Servces (ITS) Research Computing team and has served as the university’s MGHPCC lead since the consortium’s inception in 2010. “ITS is committed to helping our researchers by providing leading-edge technology resources and support. The university’s investment in Discovery and MGHPCC recognizes the importance of effective computing in identifying and solving the new research problems facing our world. I look forward to more opportunities for us to collaborate with and support our researchers.”
Established in 2012, the Discovery Cluster has steadily grown in size and breadth of services. Discovery currently offers more than 10,000+ Intel compute cores, 50TB of RAM, 2.5 PiB of high-speed parallel file system storage, multiple high-end GPUs, a Hadoop/Spark big data configuration, and support for sensitive data research at the CMS security level.
In addition to the shared access model, ITS offers several hosting and support options, including a ‘buy-in’ service model for researchers wishing to acquire and contribute resources to the cluster that are then fully managed by ITS and made available to them. Use of the cluster is governed by an advisory committee composed of faculty from many colleges.

Assistant professor Paul Whitford uses computational techniques to study the nuances of microscopic machines like the ribosome, which is responsible for protein synthesis - Image credit: Brooks Canaday.
Prof. Paul Whitford
Whitford Research Group
Department of Physics, Northeastern University
Discovery Cluster / Research Computing @ Northeastern
Northeastern professor creates software that can help in drug development Boston Business Journal
Under the Hood of the Ribosome News@Northeastern